 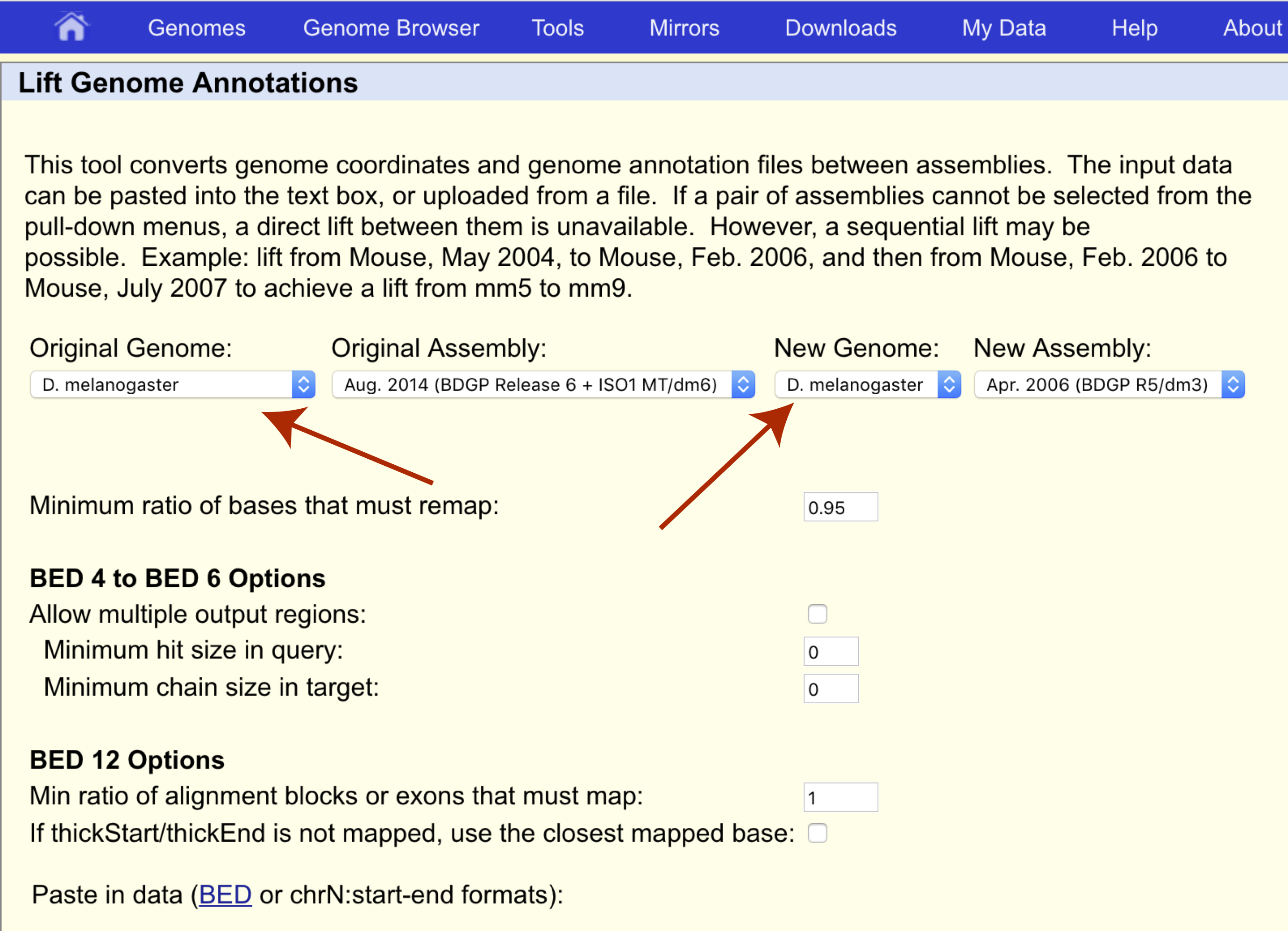
This gives you a ~633MB file: bwa pac2bwtgen hg19.fa.pac tmp.bwt & gzip tmp.bwtĪ bit-aware compression algorithm may achieve an even higher compression ratio. If you want to achieve an even smaller file size, you can compress the BWT of 2-bit file. A gzip'd 2-bit file is only ~5-10% smaller. You can compress it further with gzip, but that actually doesn't work well. For human genome, you get a file ~784MB in size. The 2-bit format typically reduces the file size down to 1/4 of its original size, unless there are too many scattered ambiguous bases. BWA does not provide utilities to convert its 2-bit representation to FASTA, either. Unlike UCSC, BWA keeps all IUB codes but loses letter cases. You can generate it separately with: bwa fa2pac -f hg19.fa UCSC's hg19 differs from GRCh37 at a few bases.īWA also produces its own 2-bit format with indexing. 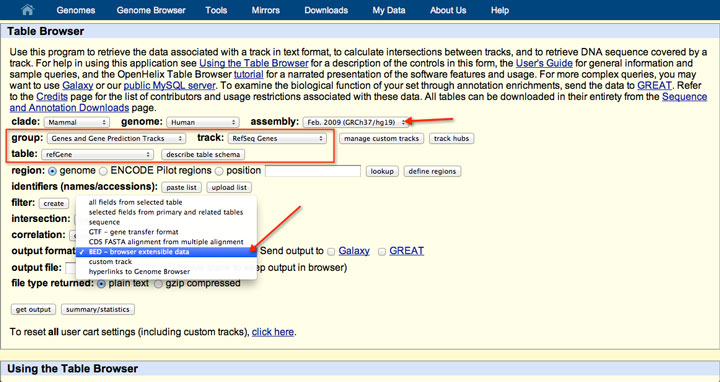
The 2-bit format loses IUB codes that GRCh37 has. These lists basically tell you that bases between offset x and y are all "N"/lowercase. As I remember, it keeps non-A/C/G/T bases and lowercases in two separate lists. This format keeps each A/C/G/T with 2 bits. For the ~3GB human genome, gzip reduces the size to ~900MB, depending on the option in use.Īnother often used format is UCSC's 2-bit format. The standard and the most common sequence format is FASTA for sure.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
